 
Rosetta homology modeling and ab initio fragment assembly with Ginzu domain predictionĬombination of ab initio folding and homology methodsĭetection of templates, alignment, modeling incl. These include alpha helices, beta strands (sheets) and reverse turns. Proprietary platform, supported on Windows, Linux and Mac Secondary structures are those repetitive structures involving H bond between amide H and carbonyl O in- the main chain. excluded ligand volumes), loop modelling, rotamer libraries for sidechain conformations, relaxation using MM forcefields. Template identification, use of multiple templates and accounting for other environments (e.g. Standalone program mainly in Fortran and Perl Satisfaction of contact and distance restraints Standalone program mainly in Fortran and Python Webserver with job manager, automatically updated fold library, genome searching and other facilities Remote template detection, alignment, 3D modeling, multi-templates, ab initio Template detection, alignment, 3D modeling Wraps external programs into automated workflowīLAST search, T-Coffee alignment, and MODELLER construction 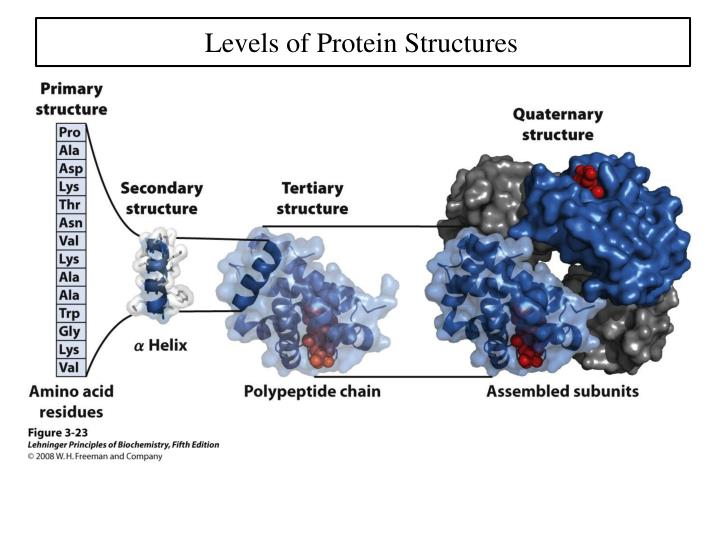 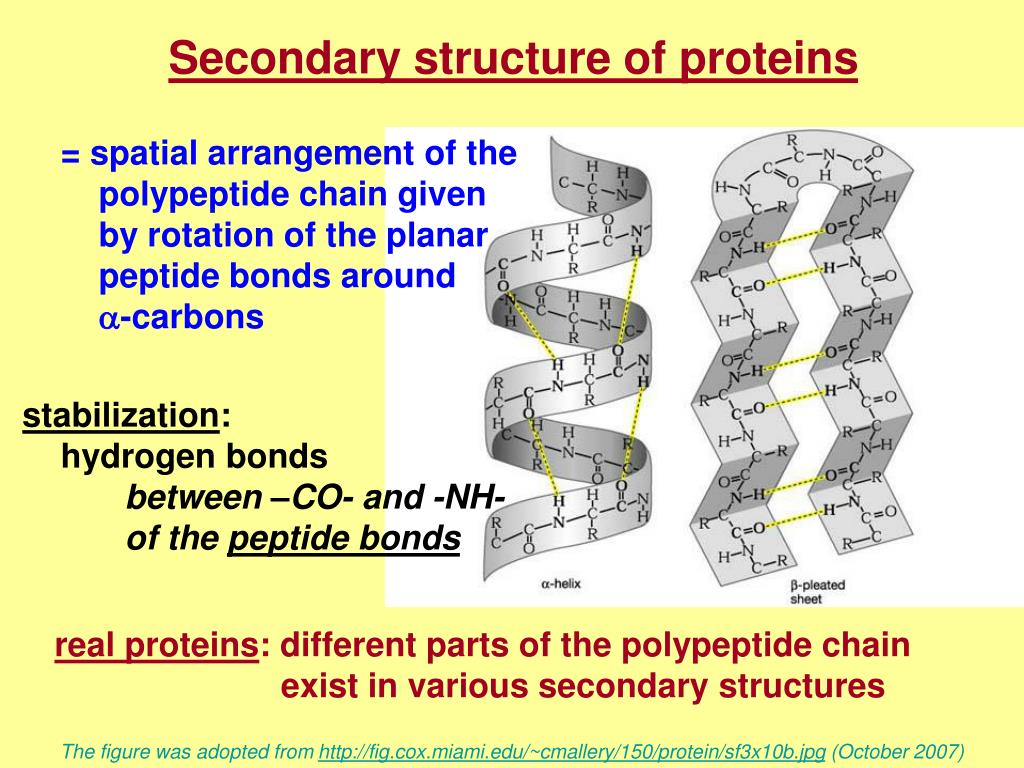
Remote homology detection, protein 3D modeling, binding site predictionĪutomated webserver and Downloadable program Secondary protein structure Tertiary structure Quaternary protein Protein. A unified interface for: Tertiary structure prediction/3D modelling, 3D model quality assessment, Intrinsic disorder prediction, Domain prediction, Prediction of protein-ligand binding residuesĪutomated webserver and some downloadable programs Proteins are made up of a long chain of amino acids. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
